The problem:
It is thought that nearly half of infertility cases have a genetic basis. Despite extensive knowledge gained from gene knockouts in mice, the genetic causes for the vast majority of idiopathic human infertilities are unknown. Traditional methods for studying inheritance, such as GWAS or linkage analysis, have been confounded by heterogeneity of infertility phenotypes and many hundreds of genes involved in reproduction. This web site summarizes the results from a project funded by the National Institute of Child Health and Human Development (R01HD082568), entitled “Identification and validation of human infertility alleles.” This project employs unique approach to identifying infertility genes in the population. The Schimenti lab, in collaboration with the lab of Haiyuan Yu at Cornell, are identifying nonsynonymous single nucleotide polymorphisms (nsSNPs) that functionally disrupt gametogenesis and cause fertility defects.
The specific aims are to:
- Use computational approaches and high-throughput in vitro assays to identify nsSNPs in human infertility genes that are likely to disrupt protein function. These alleles are then precisely modeled in mice using CRISPR/Cas genome editing.
- Phenotype the mouse models to identify those SNPs which impact gametogenesis and fertility. An overview of the project’s flow is diagrammed below:
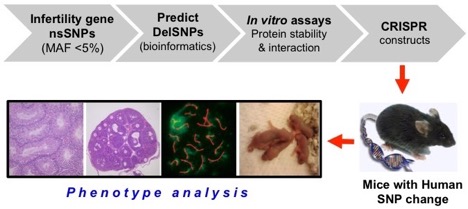
Overall, several dozen mouse models corresponding to human nsSNPs in known reproduction genes are being made and characterized. The results of our experiments are presented here in a searchable database of experimentally-validated benign and deleterious SNPs that we’ve modeled. Since the SNPs being modeled are segregating in the population, this will serve as a permanent resource for the human reproductive genetics field in the era of personalized medicine, providing solid experimental evidence for the consequences of these nsSNPs upon gene function.